MutMap-Gap: whole-genome resequencing of mutant F2 progeny bulk combined with de novo assembly of gap regions identifies the rice blast resistance gene Pii. | Semantic Scholar

Assembling the Setaria italica L. Beauv. genome into nine chromosomes and insights into regions affecting growth and drought tolerance | Scientific Reports

Scaffolding and completing genome assemblies in real-time with nanopore sequencing | Nature Communications

The Projector 2 procedure. From top to bottom: I , single (left) or... | Download Scientific Diagram

Figure 1 from GMcloser: closing gaps in assemblies accurately with a likelihood-based selection of contig or long-read alignments | Semantic Scholar

PGcloser: Fast Parallel Gap-Closing Tool Using Long-Reads or Contigs to Fill Gaps in Genomes | Semantic Scholar

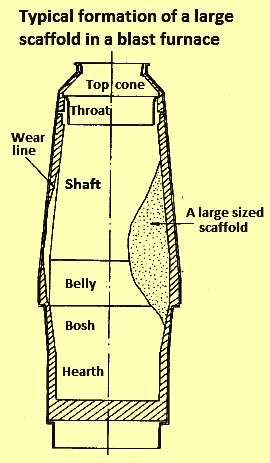
Behavior of Chromium, Nickel, Lead, Zinc, Cadmium, and Mercury in the Blast Furnace—A Critical Review of Literature Data and Plant Investigations | Industrial & Engineering Chemistry Research

A more detailed visualization of the gapFinisher pipeline workflow. a)... | Download Scientific Diagram
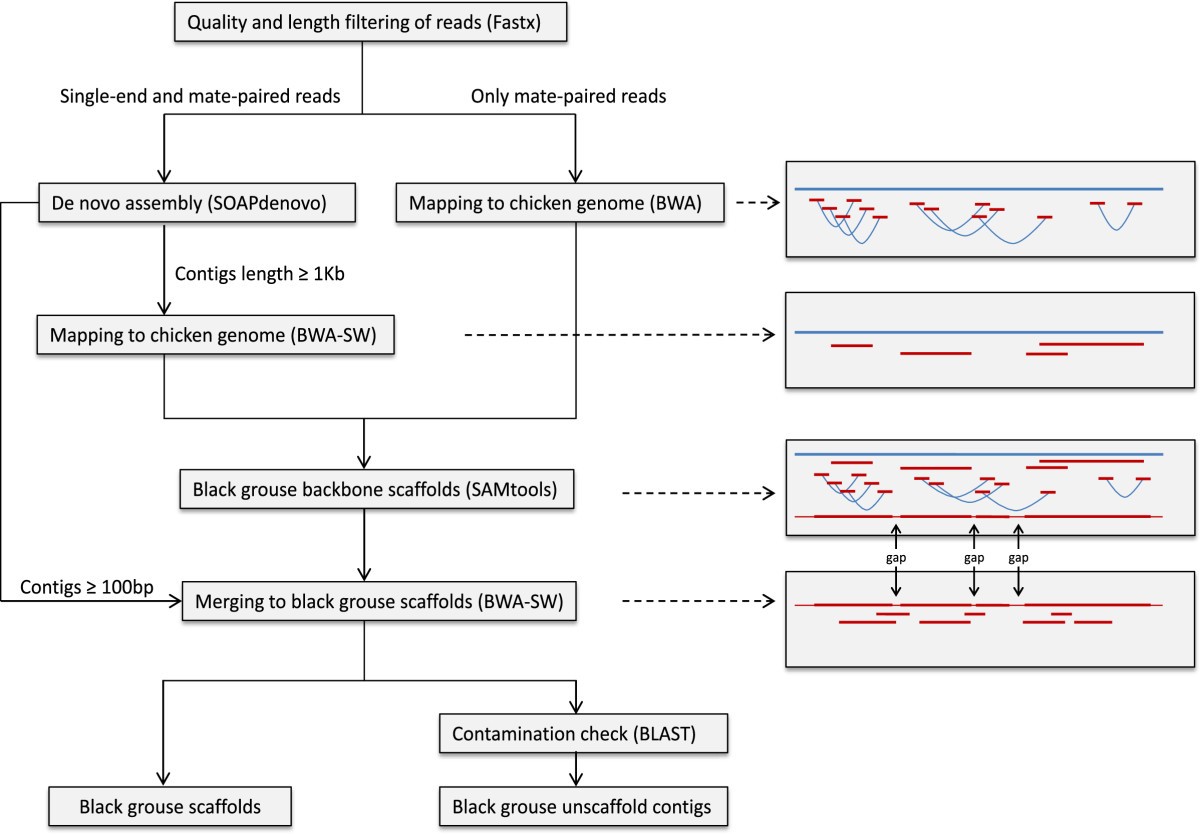
Whole genome sequencing of the black grouse (Tetrao tetrix): reference guided assembly suggests faster-Z and MHC evolution | BMC Genomics | Full Text
Semi-Automatic In Silico Gap Closure Enabled De Novo Assembly of Two Dehalobacter Genomes from Metagenomic Data | PLOS ONE
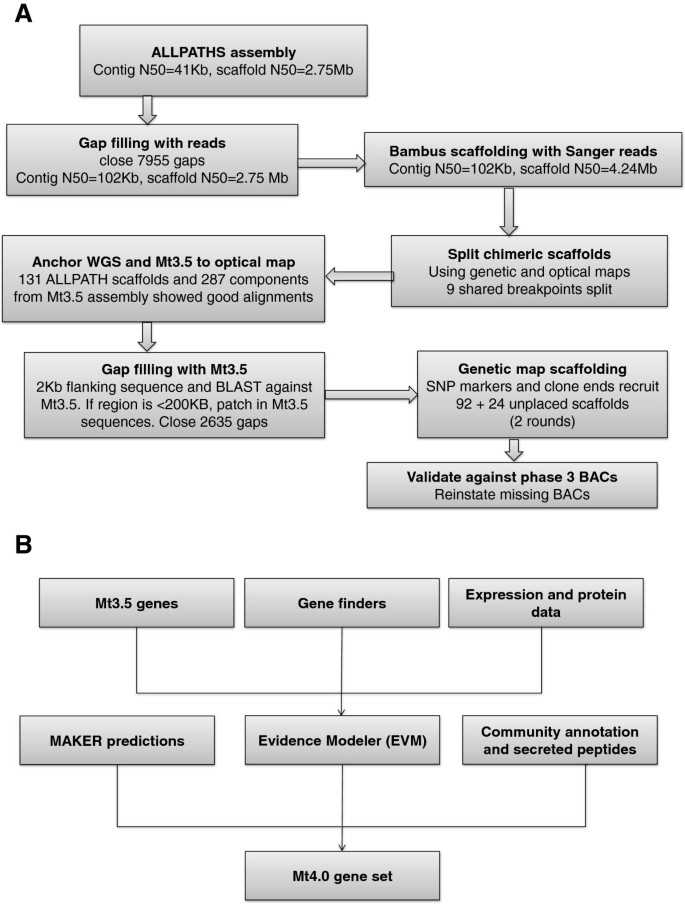
An improved genome release (version Mt4.0) for the model legume Medicago truncatula | BMC Genomics | Full Text
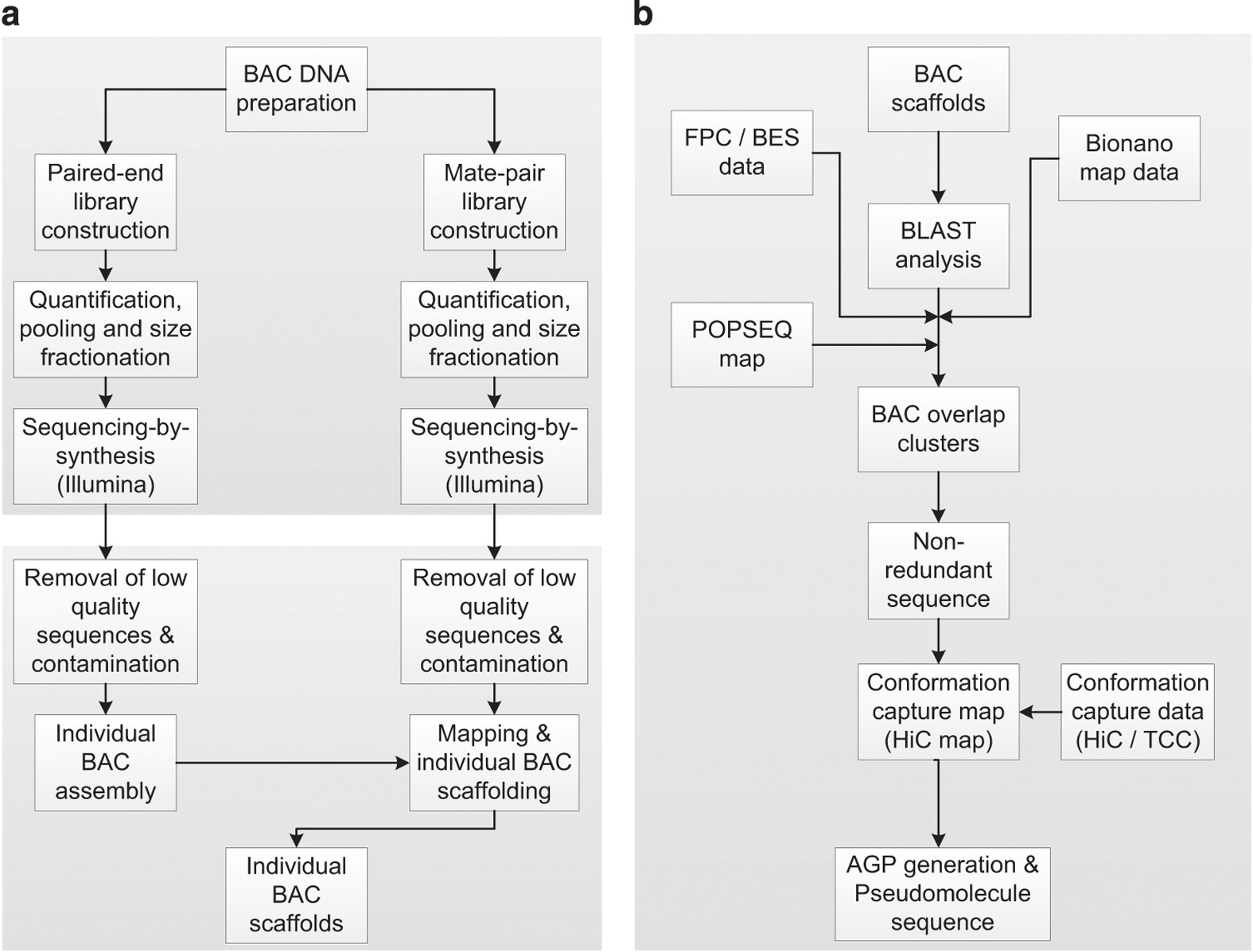




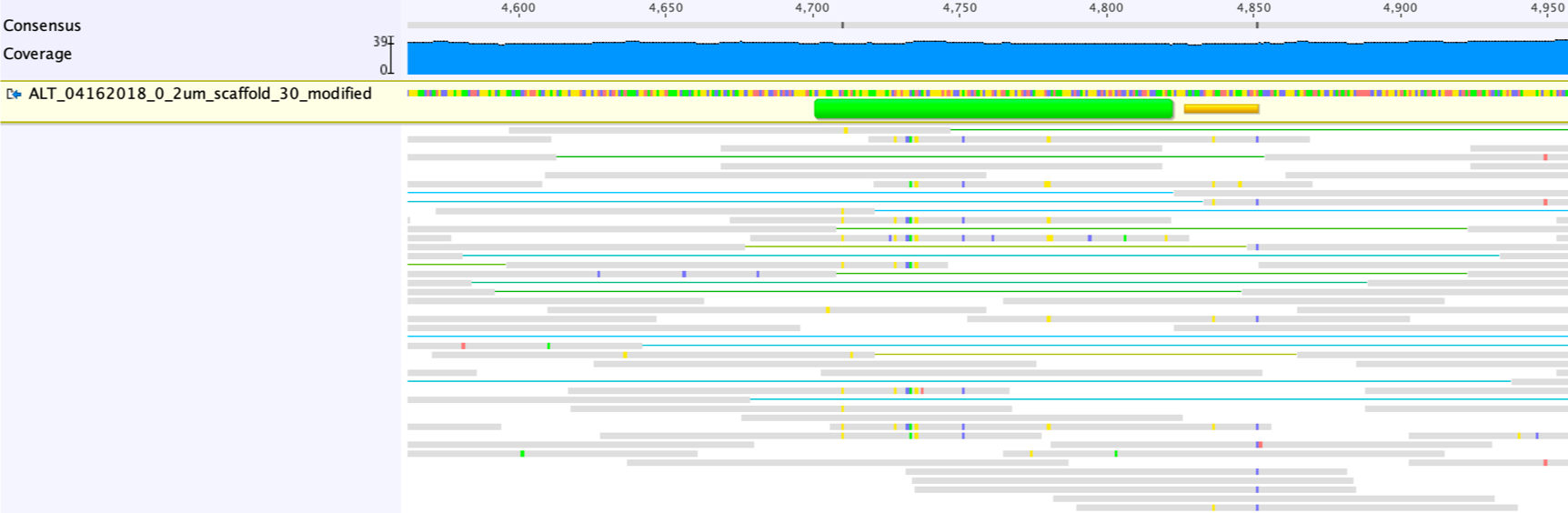
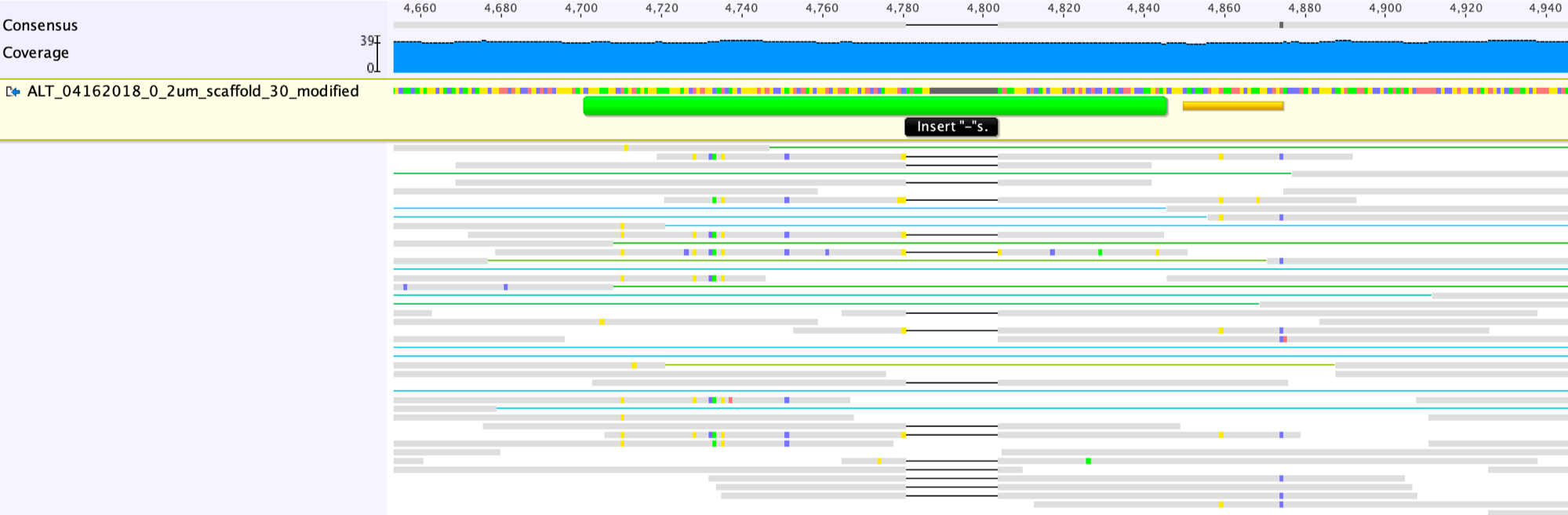
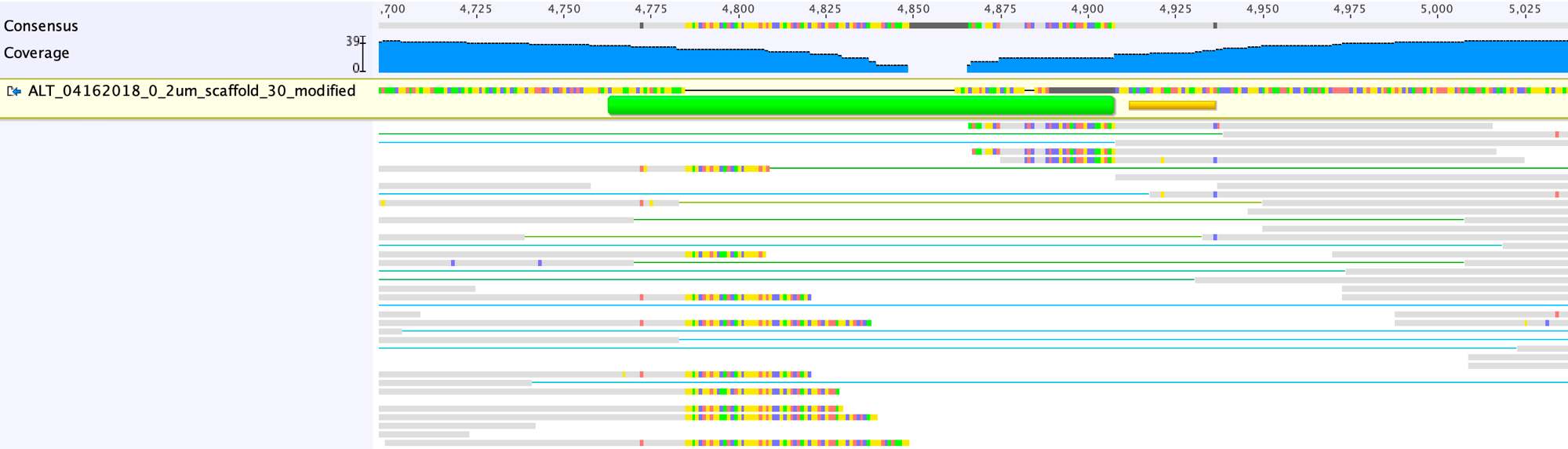
![RFfiller: a robust and fast statistical algorithm for gap filling in draft genomes [PeerJ] RFfiller: a robust and fast statistical algorithm for gap filling in draft genomes [PeerJ]](https://dfzljdn9uc3pi.cloudfront.net/2022/14186/1/fig-1-2x.jpg)